Chromosome segregation is critical to ensure that a complete copy of the parent's genetic information is passed on to each daughter cell. In most bacteria and plasmids, the correct segregation of chromosomes relies on the ParABS system, which consists of the ATPase protein ParA, the DNA-binding protein ParB and its target sequence parS. The formation of the ParB-DNA nucleoprotein complex is the key to its subsequent chromosome segregation. It typically begins with the nucleation of dimeric ParB molecules at the parS site and driven by CTP, dimeric ParB diffuses to adjacent DNA. The assembly of higher-order ParB-DNA nucleoprotein complexes requires multiple ParB dimers to aggregate to form co-condensates with DNA. However, what remained unknown was how hundreds of parS-centered ParB dimers efficiently assemble into condensates for chromosomal DNA organization and compression.
A research team led by SLST Associate Professor Sun Bo has now revealed that ParB orchestrates DNA assembly via multimerization. On July 29, this work was published inNucleic Acids Research in an article entitled “Stochastically multimerized ParB orchestrates DNA assembly as unveiled by single-molecule analysis”.
In their study, by using a single-molecule approach, the researchers detected the real-time dynamic interplay between ParB and DNA. They found that ParB dimers can spontaneously multimerize on and off DNA. The presence of CTP triggers the diffusion of ParB multimers on DNA. Furthermore, ParB multimers have the unique features of bridging and transporting DNA molecules, which are able to recognize parS. This work provides the important molecular basis for understanding how the ParABS system mediates chromosome segregation.
Second year Ph.D. candidate Guo Lijuan and first year Ph.D. candidate Zhao Yilin of Sun Bo’s group are the co-first authors of the paper. Sun Bo is the corresponding author, and ShanghaiTech is the first affiliated institute.
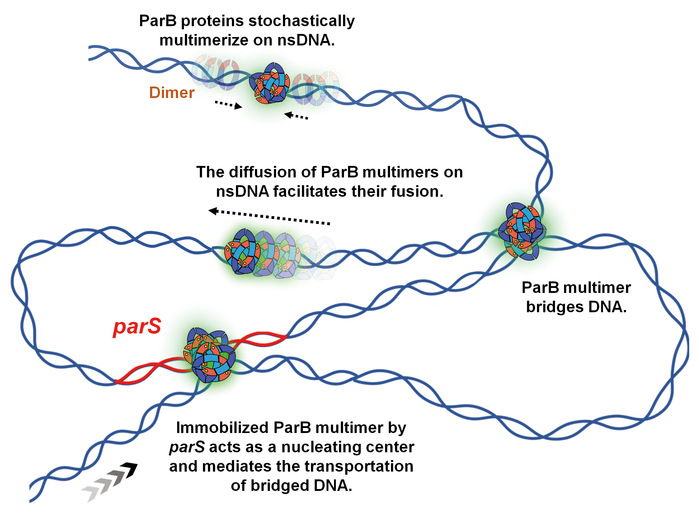
Model of ParB multimer organizing DNA assembly

