Recently, the research team led by SLST Associate Professor Sun Bo published a research article in PNAS entitled “The convergence of head-on DNA unwinding forks induces helicase oligomerization and activity transition”, revealing a novel activity transition pathway of a DNA-repair-involved helicase.
Helicases are ubiquitous molecular motors found in all organisms from phages to humans. These enzymes typically use the chemical energy from nucleoside triphosphate (NTP) binding and hydrolysis to catalyze the strand separation of the duplex form of nucleic acids and participate in almost every aspect of nucleic acid metabolisms. Moreover, increasing evidence suggests that DNA helicases are multifunctional enzymes, and besides DNA unwinding, they play additional roles in DNA replication, repair and recombination, such as protein displacement from DNA and structural rearrangement of protein complexes. For example, as one of the highly conserved RecQ family enzymes, Bloom syndrome helicase (BLM) is a model helicase that has been proven to unwind duplex DNA in its monomeric form. The BLM helicase can process other distinctive DNA substrates, recruit and displace partner proteins, and stimulate and suppress their enzymatic activities, acting as a multifunctional “caretaker” for genome stability. In addition to the monomer state, BLM can also form other oligomeric species, such as dimer and hexamer. Whether there is a correlation between its oligomeric state and cellular function and how helicases effectively perform functional switching still remain enigmatic.
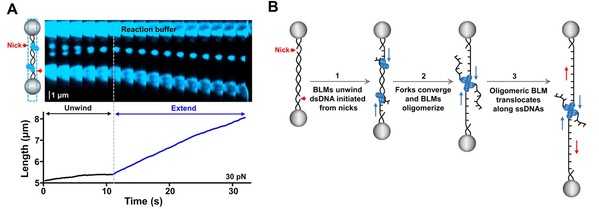
In this study, researchers employed a combined single-molecule approach. By examining the head-on collision of two BLM-mediated DNA unwinding forks, researchers found that two groups of BLM helicases, upon fork convergence, promptly oligomerize across the fork junctions. Strikingly, onsite BLM oligomerization gives rise to an immediate transition of their helicases activities from unwinding double-stranded DNA to translocating along single-stranded DNA, thus allowing for the efficient displacement of single-stranded DNA binding proteins (RPA and RAD51). These findings uncover a novel functional switch pathway of helicases and help to explain how BLM plays both pro- and anti-recombination roles in the maintenance of genome stability.
The work was carried out with the technical support from the Molecular and Cell Biology Core Facility (MCBCF) at SLST. This work was supported by the National Natural Science Foundation of China, the Natural Science Foundation of Shanghai, and ShanghaiTech University Startup funding.