SIST Associate Professor Zheng Jie, together with his collaborators, proposed a novel AI model named KG4SL, based on knowledge graph (KG) and graph neural network, and made a breakthrough in the prediction of genetic connections of Synthetic Lethality (SL). It can accelerate the discovery of anti-cancer drug targets and advance the development of AI pharmaceutical technology. This work was accepted as a regular paper, entitled “KG4SL: Knowledge Graph Neural Network for Synthetic Lethality Prediction in Human Cancers”, in the Proceedings of ISMB/ECCB (the 29th Conference on Intelligent Systems for Molecular Biology and 20th European Conference on Computational Biology). This paper was also published online in Bioinformatics.
Complex biological systems are dependent on gene-gene interactions, of which SL is a special type. Two genes are said to have the SL relationship with each other if they satisfy the following condition: when both genes are inactivated, the cell will die; but when only one of the genes is inactivated, the cell’s viability will not be affected. Therefore, SL gene pairs are potential anti-cancer drug targets for the reason that cancer cells normally have many genes mutated and inactivated, and a drug that inactivates an SL partner gene with cancer-specific mutation can selectively kill cancer cells but spare normal cells.
However, wet-lab screenings of SL pairs are afflicted with limitations such as high cost, batch effect, and off-target issue. Existing computational methods for SL prediction tend to ignore shared biological mechanisms underlying different SL pairs. To address these issues, Prof. Zheng’s group proposed a novel graph neural network (GNN)-based model of KG4SL with the involvement of a KG to capture the common biological mechanisms behind different SL gene pairs, and thereby achieve a better prediction performance and better interpretability of biological meaning.
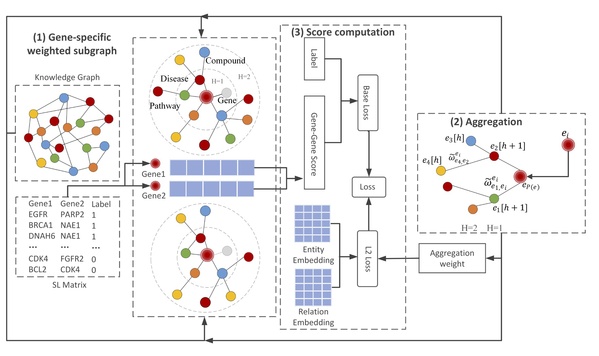
Figure 1. The framework of KG4SL. It is composed of three modules, namely Gene-specific weighted subgraph, Aggregation and Score computation. (1) Gene-specific weighted subgraph: For each SL gene pair, it constructs weighted subgraphs from the KG. (2) Aggregation: For each SL pair, it selects entities and relationships directly related to the nodes from the weighted subgraphs. Based on the assumption that the biological information can flow between nodes through edges, the information of indirectly connected entities and relationships are aggregated into feature representations of the genes. (3) Score computation: Applying a sigmoid function to the inner product of representations between two genes, it computes the probability of the two genes with SL relationship.
By incorporating a suitable KG into a graph neural network, KG4SL takes into account biological mechanisms about gene-gene interactions stored in the KG, and thereby overcomes the deficiencies of the existing assumption that SL gene pairs are independent from each other. The KG in KG4SL is from a comprehensive database on SL gene pairs called SynLethDB (http://synlethdb.sist.shanghaitech.edu.cn/v2/#/) which was developed by Prof. Zheng’s group. Compared with all baseline models, KG4SL achieved a significant improvement in performance. Moreover, as shown in Figure 2, KG4SL has more discriminatory power than the other three existing models, as each of the latter relies on KG or SL data alone or combines them in a naive way.
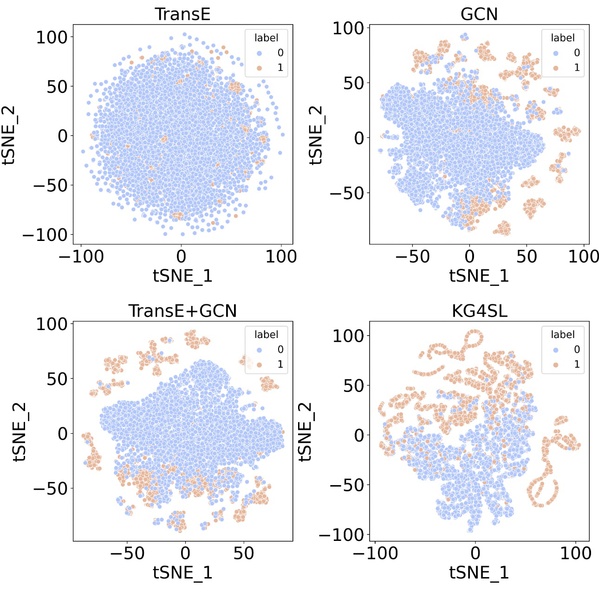
Figure 2. Visualization of SL interactions. The TransE model uses only the KG information, and the GCN model only the SL interactions for model training. The TransE+GCN model integrates the information from the KG and SL matrix. In the above plots, dot colors indicate whether SL relationship exists between a pair of genes, where SL pair is in orange and non-SL pair in blue. The plots show that KG4SL has more discriminatory power than the other models in distinguishing SL from non-SL gene pairs.
This was the first time that KG was incorporated into the task of SL prediction, and by doing so KG4SL achieved a good performance. This suggests that GNN-based deep learning models can more effectively solve complex problems in biomedical and pharmaceutical fields by integrating knowledge and data. The SL gene pairs that are newly predicted can help biologists speed up screening for new anti-cancer drug targets, thereby accelerating the progress of drug discovery using AI technology. In addition, KG4SL is promising to uncover biological mechanisms behind the SLs using KG, making deep learning models more interpretable and helpful for biological knowledge discovery.
This work was primarily completed by Prof. Zheng’s Lab from the Smart Medical Information Research Center (SMIRC) of SIST. Two first year Master students of SIST, Wang Shike and Xu Fan, are co-first authors. Dr. Yong Liu, Senior Research Scientist at Nanyang Technological University (NTU), Singapore, participated in experimental and theoretical work. Prof. Zheng Jie and Dr. Min Wu, Senior Scientist at Agency of Science and Technology Advancement and Research (A*STAR), Singapore, are the co-corresponding authors.
ISMB/ECCB is a flagship conference on Bioinformatics and Computational Biology. From among 289 manuscripts submitted to ISMB/ECCB 2021, only 55 were accepted, with a low acceptance rate of around 19%.
Link to this paper: https://academic.oup.com/bioinformatics/article/37/Supplement_1/i418/6319703

