A team of scientists led by Dr. Jia Chen and Dr. Xingxu Huang from ShanghaiTech’s School of Life Science and Technology and Dr. Li Yang from the CAS-MPG Partner Institute for Computational Biology at the Chinese Academy of Sciences, have developed a series of novel base editors (BEs) using a spectrum of cytidine deaminases from different species, including human APOBEC3A (hA3A). Their work, entitled “Efficient base editing in methylated regions with a human APOBEC3A-Cas9 fusion,” was published online in Nature Biotechnology on August 20, 2018.
Base editors (BEs) are developed by combining different nucleotide deaminase family members, including cytidine deaminase family members (e.g., the apolipoprotein B mRNA-editing enzyme, the catalytic polypeptide-like (APOBEC) or activation-induced deaminase (AID)) and adenosine deaminase family members (e.g., adenosine deaminase acting on RNA (ADAR)), with the CRISPR-Cas system (e.g., CRISPR/Cas9 and CRISPR/Cpf1). Various BEs have been used for targeted C-to-T/A-to-G base editing in different species. Numerous human diseases have been reported to be driven by point mutations in genomic DNA. With recently developed BEs, these disease-related point mutations can be corrected, providing new therapeutic options. The newly-developed base editing technology has been highlighted by Science as one of its Top 10 breakthrough[s] in 2017.
By analyzing disease-related T-to-C mutations that can be theoretically reverted to thymines by BEs, the research team found that roughly 43% of them are on cytosines in the context of CpG dinucleotides. It is well known that C of CpG is usually methylated in mammalian cells, and the methylation of C strongly suppresses the cytidine deamination activity of some APOBEC/AID deaminases. Previously developed BEs that were using rat APOBEC1 (rA1) cytidine deaminase were inefficient in editing cytosines in highly-methylated regions. To construct BEs for efficient C-to-T base editing in highly methylated regions, the researchers developed a series of BEs by fusing Cas9 nickase with a dozen of different APOBEC/AID deaminases, and then they showed that most of these BEs can be used for C-to-T base editing. Importantly, a novel BE using hA3A (hA3A-BE) and its engineered versions with narrower editing windows can mediate efficient C-to-T base editing in regions with high methylation levels, as well as in other examined regions within different contexts. Overall, these newly developed hA3A-BEs can comprehensively induce efficient base editing in all the examined contexts, including both methylated DNA regions and GpC dinucleotides.
This study was supported by grants from the National Natural Science Foundation of China, the Ministry of Science and Technology, the Shanghai Municipal Science and Technology Commission, and ShanghaiTech University.
Read more at: www.nature.com/articles/nbt.4198
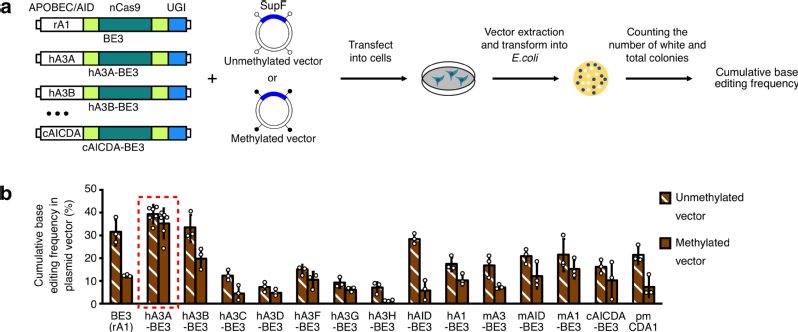
a)Construction of various BEs and screening for efficient base editing in highly methylated regions.
b)hA3A-BE3 can induce efficient base editing in highly methylated regions.